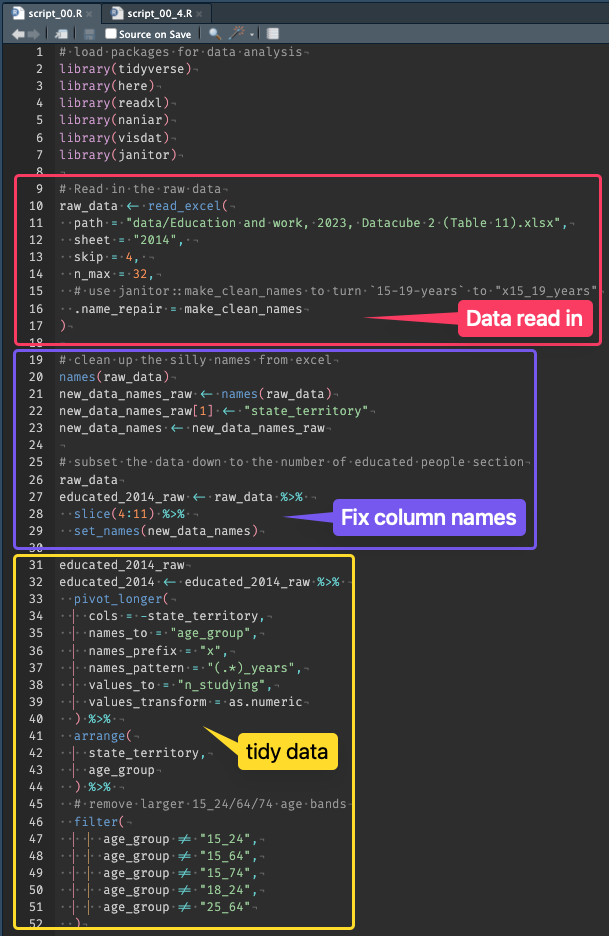
Error in max_minus_min(penguins$species): is.numeric(x) is not TRUEAdvanced Statistical Programming using R
Week 2: Scripts, Functions & Refactoring
2026-04-22
Reminders
- Register for course on module (see email)
- Bookmark course website: https://soda-lmu.github.io/StatProg2-2026-SoSe/
- Follow installation & setup requirements: https://soda-lmu.github.io/StatProg2-2026-SoSe/setup.html
- Check the updated schedule for changes due to holidays: https://soda-lmu.github.io/StatProg2-2026-SoSe/#schedule
Contact Us
For individual questions: statprog[@]stat.uni-muenchen.de For course related questions: https://moodle.lmu.de/mod/forum/view.php?id=2498223
Last Week
- Introduction to the course
- Reviewed the data science workflow
- Crash course in git & CLI
Reviewing RStudio with the Knife / Kitchen analogy
- R = the knife (the tool that actually cuts/does the work)
- RStudio = the kitchen (workspace, counters, cupboards)
- R Script = the recipe card you write ahead of time
- Console = the stove / cooking zone (where you execute the cooking)
- Terminal = talks to the kitchen manager (code outside of R, on your whole computer)
- R Markdown/Quarto = the cookbook page (with recipe + photos + notes)
- File paths = locations of ingredients (R scripts, data, images)
- absolute vs. relative
Command Line
Navigate
Create & remove
Tip
Windows uses cd for pwd, dir for ls, type for cat, and del for rm.
Git for version history
Set up & inspect
Stage & commit
The three areas
working directory → staging area → repository
(your files) (git add) (git commit)Think of it as:
- Edit files as normal
- Stage the changes you want to keep (
git add) - Commit a named snapshot (
git commit)
Why command line git?
Safety first (with agentic models)!


This Week
- Individual reflection log – git practice!
- Reviewing functions in R
- Testing functions
- Writing (better) functions (for data science tasks)
Individual Reflection Log
See “Reflection Prompts” on the course website
- The reflection log lives in the same git repository you set up in last week’s practical
- Each week you’ll answer a small set of prompts tracking what you learned around the course
- Each answer is committed as a separate commit — practising the version control workflow from day one
Tip
Find this week’s prompts on the course website under Reflection Prompts. Use the provided prompts or write your own and add to individual-reflection-log.qmd. Make sure to commit each week!
Review: Writing Functions
From StatProg 1
What are functions?
A function is a reusable block of code designed to perform a specific task. It takes input (arguments), processes it, and returns output (or performs an action).
Why use functions?
- Avoid repeating yourself (DRY)
- Easier to test and debug in isolation
- Give a name to a chunk of logic
Examples
| Function | Input | Action | Output |
|---|---|---|---|
sum(a, b) |
two numbers | a + b |
the sum |
print(x) |
any object | display it | (side effect) |
ggplot(df, aes(...)) |
data + aesthetics | build a plot | plot object |
Defining a function in R
Step 1: working interactive code
Step 2: wrap it in a function — add NO new functionality
Function syntax in R
| Part | Name | What it does |
|---|---|---|
my_function |
function name | how you call it |
function(...) |
signature | declares the function |
arg1, arg2 |
formals / parameters | inputs the function accepts |
arg2 = default |
default value | used when argument is omitted |
{ ... } |
body | the code that runs |
return(...) |
return value | what the function hands back |
Tip
If there is no return(), R returns the value of the last expression in the body.
Testing Functions
Based on: https://stat545.com/functions-part1.html
Why test functions?
- Verify the function does what you think it does — not what you hope it does
- Catch silent errors — wrong output with no error message is worse than a crash
- Build confidence to reuse the function later, in other scripts or projects
- Isolate bugs — a tested function is one less place to look when something breaks downstream
- Document intent — tests show what inputs are expected and what the output should be
Tip
You don’t need a formal testing framework to start. “Does this give the right answer on an input I can check by hand?” is already a test.
Testing Sensible Inputs
Test on inputs where you know the answer
Tip
Check by hand for exact confirmation, or compare it with data type/domain expectations (e.g. is this a reasonable height?) as a sense check.
Stress-testing functions
After testing that functions work as expected with sensible inputs, it is also useful to try and intentionally ‘break’ your functions with unexpected inputs.
Expected failures — R throws an error
Note
Are the messages helpful enough to diagnose the problem quickly?
Silent failures — R returns something, no error
Warning
R’s eagerness to make sense of your input works against you here. Can imagine similar issues in LLM-assisted workflows?
Validating Function inputs
stopifnot() — quick but blunt
Catches the problem, but the message isn’t helpful.
if / stop() — custom message
Error in max_minus_min(penguins$species): Expected numeric, got factorTemplate: “you gave me THIS, but I need THAT”
When testing is hard
max_minus_min() is easy to test because it returns a single number. Real data science functions often return something harder to check:
Complex output
What exactly do you test? The coefficients? The residuals? Whether it runs at all?
Tip
Testing something specific is better than testing nothing.
Strategies
- Test one specific, checkable property at a time
- Snapshot a known-good output and compare
- Test the pieces separately
Matters of design & style
Things to consider when writing a function…
- names, names, names!
- how much should a function do?
- what inputs should it take?
- how should inputs be accepted?
- what outputs should the function provide?
- can I make more function easier to test?
- what should the error messages say?
Naming functions (as verbs)
A function’s name should describe what it does, not what it is.
Tip
If you struggle to name a function, it may be doing too many things.
Extension: Documenting (package) functions
roxygen2 turns structured comments placed directly above a function into ?help documentation when packaged.
Core tags
| Tag | What it documents |
|---|---|
@title |
one-line title (or first line) |
@description |
longer description |
@param name |
an argument |
@return |
what the function returns |
@examples |
runnable example code |
@export |
marks function as public in a package |
Example
#' Range of a numeric vector
#'
#' Returns max minus min of a numeric vector,
#' ignoring missing values.
#'
#' @param x A numeric vector.
#' @return A single numeric value.
#' @examples
#' max_minus_min(1:10)
#' max_minus_min(c(3, 1, 4, 1, 5, 9))
max_minus_min <- function(x) {
if (!is.numeric(x))
stop("Expected numeric, got ", class(x)[1])
max(x, na.rm = TRUE) - min(x, na.rm = TRUE)
}Writing better error messages
- give a concise but informative problem statement
- error location & details
- provide solution hints
Tip
Thinking about error messages can also help you find silent errors or possible breakage points in your functions
Extension: Better errors with cli
cli_abort() is used throughout the tidyverse — you will have seen its output already.
Error in `max_minus_min()`:
! `x` must be a numeric vector.
✖ You supplied a <factor>.| Token | Renders as |
|---|---|
{.arg x} |
styled argument name: `x` |
{.cls {class(x)}} |
styled class name: <factor> |
"x" = "..." |
bullet prefixed with ✖ |
Avoid writing greedy functions
It’s easy to write one big function that does everything:
analyse_penguins <- function(df) {
df <- df |> filter(!is.na(bill_length_mm))
summary_tbl <- df |> group_by(species) |>
summarise(mean_bill = mean(bill_length_mm), n = n())
p <- ggplot(df, aes(bill_length_mm, flipper_length_mm, colour = species)) +
geom_point()
list(summary = summary_tbl, plot = p)
}Better: break it into focused, testable pieces
Each piece is easier to name, test, and reuse independently.
What granularity? Task abstractions
The right scope depends on the task. Consider asking an LLM to write code for these tasks:
| Task Prompt | How many functions needed? |
|---|---|
| “analyse the data” | unclear — could be anything |
| “run a linear regression” | 1–2, but what inputs and outputs? |
| “regress bill on flipper by species” | 1 function, clear inputs |
| “return the slope and p-value from a regression” | 1 function, specific output |
What granularity? Task abstractions
The right scope depends on the task. Consider asking an LLM to write code for these tasks:
| Task Prompt | How many functions needed? |
|---|---|
| “analyse the data” | unclear — could be anything |
| “run a linear regression” | 1–2, but what inputs and outputs? |
| “regress bill on flipper by species” | 1 function, clear column inputs |
| “return the slope and p-value from a regression” | 1 function, specific output |
Specific and modular!
More specific task descriptions produce better functions — whether you’re writing them yourself or prompting an LLM.
Writing (better) functions
Based on:
Why write (better) functions?
Well written functions help us:
- Manage complexity via abstraction
- Make code easier to reason about (for others and future you!)
- Express intentions, processes and decisions (within a data analysis context)
Moving beyond DRY (Don’t Repeat Yourself)
DRY — the classic rule:
If you copy-paste code 3 times, write a function.
Targets repetition in the text of your code.
DRRY — the extension:
If you re-read code 3 times, write (or improve) a function.
Targets cognitive load — the effort of understanding your own code.
Tip
Re-reading to spot a difference, to debug, or to remember what a block does are all signals that a function (with a good name) would help.
Writing DRRY functions
Two approaches to designing better functions
Outside-In: Interface design
- Write the function call before you write the body
- Imagine the function already exists.
- Think about the where and when you would call this function
- Fill in the inside of the function
Inside-Out: Refactoring
- Start with working interactive code
- Don’t just wrap in a monster function!
- Identify chunks of code that express one task or idea
- Give these chunks sensible names!
- Put these chunks into functions
- Go back outside and think about how these functions work together
Outside to Inside and Back again
The outside (interface) shapes the inside (implementation) — and vice versa. Designing the call first surfaces decisions you would otherwise hit mid-implementation.
Outside-in: Principles
Write the call before you write the body — imagine the function already exists.
Ask three questions:
| Question | Example | |
|---|---|---|
| Task | What am I trying to do? What evokes the action? | Is X different when Y is missing? |
| Inputs | What do I need to provide? | the variable going missing; the variable affected |
| Output | What am I returning? | a t-test result |
Tip
Once the interface feels right, writing the body is easier.
Outside-in: Context
To answer the questions of task, inputs and outputs, consider where in your data analysis this function will exist?
- What data tasks are you doing before (e.g. preparing tidy data)
- What inputs are needed and available to your function (a table? model parameters?)
- What should this function pass on to other parts of the data analysis?
Outside-in: Naming
Good names take a few tries — that is expected, not a sign something is wrong.
Tip
The appropriateness of names is subjective and context dependent! Use the DRRY principle of reducing cognitive load to guide you (but know there are no hard and fast rules!).
Inside-Out: Breaking code into chunks
50 lines of code
Is not 50 ideas
Chunk code into ideas
Reason with them
Find the complexity
Abstract complexity
Beware! Abstractions all the way down: ggplot2
Ways to reuse ggplot code
Custom ggplot2 code often involves:
- data preparation
- aesthetic mappings
- multiple custom layers (
geom,coord,scaleetc.)
Given working ggplot2 code (insides), how might we package this into one or more reusable functions?
Copy & Paste
Store layers as objects or in lists
💡 Useful if components are exactly the same each time!
Store layers as objects or in lists
💡 Useful if components are exactly the same each time!
Store layers as objects or in functions
Helper functions: Convenience vs. flexibility
What about customisation?
ggweekly::ggweek_planner(
start_day = lubridate::today(),
end_day = start_day +
lubridate::weeks(8) - lubridate::days(1),
highlight_days = NULL,
week_start = c("isoweek", "epiweek"),
week_start_label = c("month day", "week", "none"),
show_day_numbers = TRUE,
show_month_start_day = TRUE,
show_month_boundaries = TRUE,
highlight_text_size = 2,
month_text_size = 4,
day_number_text_size = 2,
month_color = "#f78154",
day_number_color = "grey80",
weekend_fill = "#f8f8f8",
holidays = ggweekly::us_federal_holidays,
font_base_family = "PT Sans",
font_label_family = "PT Sans Narrow",
font_label_text = NULL
)Similar function, different wrapper?
- how many arguments does
ggcal()have? - what format should
dates,fillsbe?
Another helpful calendar maker?
calendR(
year = format(Sys.Date(), "%Y"),
month = NULL,
from = NULL,
to = NULL,
start = c("S", "M"),
orientation = c("portrait", "landscape"),
title,
title.size = 20,
title.col = "gray30",
subtitle = "",
subtitle.size = 10,
subtitle.col = "gray30",
text = NULL,
text.pos = NULL,
text.size = 4,
text.col = "gray30",
special.days = NULL,
special.col = "gray90",
gradient = FALSE,
low.col = "white",
col = "gray30",
lwd = 0.5,
lty = 1,
font.family = "sans",
font.style = "plain",
day.size = 3,
days.col = "gray30",
weeknames,
weeknames.col = "gray30",
weeknames.size = 4.5,
week.number = FALSE,
week.number.col = "gray30",
week.number.size = 8,
monthnames,
months.size = 10,
months.col = "gray30",
months.pos = 0.5,
mbg.col = "white",
legend.pos = "none",
legend.title = "",
bg.col = "white",
bg.img = "",
margin = 1,
ncol,
lunar = FALSE,
lunar.col = "gray60",
lunar.size = 7,
pdf = FALSE,
doc_name = "",
papersize = "A4"
)- Is this using ggplot2?
- What are the inputs?
- Internal data preparation?
- …?
ASIDE: Inspecting Functions
To look at the defintions of your own custom functions:
To look at functions from packages:
Over-sized functions?
function (year = format(Sys.Date(), "%Y"), month = NULL, from = NULL,
to = NULL, start = c("S", "M"), orientation = c("portrait",
"landscape"), title, title.size = 20, title.col = "gray30",
subtitle = "", subtitle.size = 10, subtitle.col = "gray30",
text = NULL, text.pos = NULL, text.size = 4, text.col = "gray30",
special.days = NULL, special.col = "gray90", gradient = FALSE,
low.col = "white", col = "gray30", lwd = 0.5, lty = 1, font.family = "sans",
font.style = "plain", day.size = 3, days.col = "gray30",
weeknames, weeknames.col = "gray30", weeknames.size = 4.5,
week.number = FALSE, week.number.col = "gray30", week.number.size = 8,
monthnames, months.size = 10, months.col = "gray30", months.pos = 0.5,
mbg.col = "white", legend.pos = "none", legend.title = "",
bg.col = "white", bg.img = "", margin = 1, ncol, lunar = FALSE,
lunar.col = "gray60", lunar.size = 7, pdf = FALSE, doc_name = "",
papersize = "A4")
{
if (year < 0) {
stop("You must be kidding. You don't need a calendar of a year Before Christ :)")
}
wend <- TRUE
l <- TRUE
if ((!is.null(from) & is.null(to))) {
stop("Provide an end date with the 'to' argument")
}
if ((is.null(from) & !is.null(to))) {
stop("Provide a start date with the 'from' argument")
}
if (is.character(special.days) & length(unique(na.omit(special.days))) !=
length(special.col)) {
stop("The number of colors supplied on 'special.col' argument must be the same of length(unique(na.omit(special.days)))")
}
if (length(unique(start)) != 1) {
start <- "S"
}
if (length(unique(orientation)) != 1) {
orientation <- "landscape"
}
if (missing(ncol)) {
ncol <- ifelse(orientation == "landscape" | orientation ==
"l", 4, 3)
}
match.arg(start, c("S", "M"))
match.arg(orientation, c("landscape", "portrait", "l", "p"))
match.arg(papersize, c("A6", "A5", "A4", "A3", "A2", "A1",
"A0"))
if (!is.null(month)) {
if (month > 12) {
stop("There are no more than 12 months in a year")
}
if (month <= 0) {
stop("Months must be between 1 and 12")
}
if (is.character(month)) {
stop("You must provide a month in a numeric format, between 1 and 12")
}
}
months <- format(seq(as.Date("2016-01-01"), as.Date("2016-12-01"),
by = "1 month"), "%B")
if (!is.null(text) && is.null(text.pos)) {
warning("Select the number of days for the text with the 'text.pos' argument")
}
if (is.null(text) && !is.null(text.pos)) {
warning("Add text with the 'text' argument")
}
if (missing(weeknames)) {
up <- function(x) {
substr(x, 1, 1) <- toupper(substr(x, 1, 1))
x
}
Day <- seq(as.Date("2020-08-23"), by = 1, len = 7)
weeknames <- c(up(weekdays(Day))[2:7], up(weekdays(Day))[1])
}
if (!is.null(from) & !is.null(to)) {
if (as.numeric(as.Date(from) - as.Date(to)) > 0) {
stop("'to' must be posterior to 'from'")
}
if (lunar == TRUE) {
l <- FALSE
warning("Lunar phases are only available for monthly calendars")
}
mindate <- as.Date(from)
maxdate <- as.Date(to)
weeknames <- substring(weeknames, 1, 3)
}
else {
if (is.null(month)) {
mindate <- as.Date(format(as.Date(paste0(year, "-0",
1, "-01")), "%Y-%m-01"))
maxdate <- as.Date(format(as.Date(paste0(year, "-12-",
31)), "%Y-%m-31"))
weeknames <- substring(weeknames, 1, 3)
}
else {
if (month >= 10) {
mindate <- as.Date(format(as.Date(paste0(year,
"-", month, "-01")), "%Y-%m-01"))
}
else {
mindate <- as.Date(format(as.Date(paste0(year,
"-0", month, "-01")), "%Y-%m-01"))
}
maxdate <- seq(mindate, length = 2, by = "months")[2] -
1
}
}
if (!is.null(from) & !is.null(to)) {
if (as.Date(to) - as.Date(from) > 366) {
stop("'from' and 'to' can't me more than 1 year appart")
}
if (as.numeric(as.Date(to) - as.Date(from)) > 0) {
filler <- dplyr::tibble(date = seq(mindate, maxdate,
by = "1 day"))
dates <- seq(mindate, maxdate, by = "1 day")
}
else {
stop("'to' must be posterior to 'from'")
}
}
else {
filler <- dplyr::tibble(date = seq(mindate, maxdate,
by = "1 day"))
dates <- seq(mindate, maxdate, by = "1 day")
}
fills <- numeric(length(dates))
texts <- character(length(dates))
texts[text.pos] <- text
moon_m <- suncalc::getMoonIllumination(date = dates, keep = c("fraction",
"phase", "angle"))
moon <- moon_m[, 2]
right <- ifelse(moon_m[, 4] < 0, TRUE, FALSE)
if (is.character(special.days)) {
if (length(special.days) != length(dates)) {
if (special.days != "weekend") {
stop("special.days must be a numeric vector, a character vector of the length of the number of days of the year or month or 'weekend'")
}
else {
wend <- FALSE
}
}
if (gradient == TRUE) {
warning("Gradient won't be created as 'special.days' is of type character. Set gradient = FALSE in this scenario to avoid this warning")
if (legend.title != "" & legend.pos == "none") {
warning("Legend title specified, but legend.pos == 'none', so no legend will be plotted")
}
}
else {
if (length(special.days) != length(dates) & (legend.pos !=
"none" | legend.title != "")) {
legend.pos = "none"
warning("gradient = FALSE, so no legend will be plotted")
}
}
}
else {
if (gradient == FALSE) {
if (length(special.days) != length(dates) & (legend.pos !=
"none" | legend.title != "")) {
legend.pos = "none"
warning("gradient = FALSE, so no legend will be plotted")
}
}
else {
if (legend.title != "" & legend.pos == "none") {
warning("Legend title specified, but legend.pos == 'none', so no legend will be plotted")
}
}
if (gradient == TRUE & (length(special.days) != length(dates))) {
stop("If gradient = TRUE, the length of 'special.days' must be the same as the number of days of the corresponding month or year")
}
}
if (start == "M") {
weekdays <- weeknames
t1 <- dplyr::tibble(date = dates, fill = fills) %>% right_join(filler,
by = "date") %>% mutate(dow = ifelse(as.numeric(format(date,
"%w")) == 0, 6, as.numeric(format(date, "%w")) -
1)) %>% mutate(month = format(date, "%B")) %>% mutate(woy = as.numeric(format(date,
"%W"))) %>% mutate(year = as.numeric(format(date,
"%Y"))) %>% mutate(month = toupper(factor(month,
levels = months, ordered = TRUE))) %>% mutate(monlabel = month)
if (!is.null(month)) {
t1$monlabel <- paste(t1$month, t1$year)
}
t2 <- t1 %>% mutate(monlabel = factor(monlabel, ordered = TRUE)) %>%
mutate(monlabel = fct_inorder(monlabel)) %>% mutate(monthweek = woy -
min(woy), y = max(monthweek) - monthweek + 1) %>%
mutate(weekend = ifelse(dow == 6 | dow == 5, 1, 0))
if (all(special.days == 0) == TRUE || length(special.days) ==
0) {
special.col <- "white"
}
else {
if (is.character(special.days)) {
if (length(special.days) == length(dates)) {
fills <- special.days
}
else {
if (special.days == "weekend") {
fills <- t2$weekend
}
}
}
else {
if (gradient == TRUE) {
fills <- special.days
}
else {
fills[special.days] <- 1
}
}
}
}
else {
weekdays <- c(weeknames[7], weeknames[1:6])
t1 <- dplyr::tibble(date = dates, fill = fills) %>% right_join(filler,
by = "date") %>% mutate(dow = as.numeric(format(date,
"%w"))) %>% mutate(month = format(date, "%B")) %>%
mutate(woy = as.numeric(format(date, "%U"))) %>%
mutate(year = as.numeric(format(date, "%Y"))) %>%
mutate(month = toupper(factor(month, levels = months,
ordered = TRUE))) %>% mutate(monlabel = month)
if (!is.null(month)) {
t1$monlabel <- paste(t1$month, t1$year)
}
t2 <- t1 %>% mutate(monlabel = factor(monlabel, ordered = TRUE)) %>%
mutate(monlabel = fct_inorder(monlabel)) %>% mutate(monthweek = woy -
min(woy), y = max(monthweek) - monthweek + 1) %>%
mutate(weekend = ifelse(dow == 0 | dow == 6, 1, 0))
if (all(special.days == 0) == TRUE || length(special.days) ==
0) {
special.col <- "white"
}
else {
if (is.character(special.days)) {
if (length(special.days) == length(dates)) {
fills <- special.days
}
else {
if (special.days == "weekend") {
fills <- t2$weekend
}
}
}
else {
if (gradient == TRUE) {
fills <- special.days
}
else {
fills[special.days] <- 1
}
}
}
}
df <- data.frame(week = weekdays, pos.x = 0:6, pos.y = rep(max(t2$monthweek) +
1.75, 7))
if (missing(title)) {
if (!is.null(from) & !is.null(to)) {
title <- paste0(format(as.Date(from), "%m"), "/",
format(as.Date(from), "%Y"), " - ", format(as.Date(to),
"%m"), "/", format(as.Date(to), "%Y"))
}
else {
if (is.null(month)) {
title <- year
}
else {
title <- levels(t2$monlabel)
}
}
}
if (week.number == FALSE) {
week.number.col <- "transparent"
}
if (is.null(month) | (!is.null(from) & !is.null(to))) {
if (!missing(monthnames)) {
if (length(monthnames) == length(levels(t2$monlabel))) {
t2$monlabel <- factor(t2$monlabel, labels = monthnames)
}
else {
stop("The length of 'monthname's must equal to the number months")
}
}
if (lunar == TRUE & l != FALSE) {
warning("Lunar phases are only available for monthly calendars")
}
if (gradient == TRUE || !missing(special.days)) {
p <- ggplot(t2, aes(dow, woy + 1)) + geom_tile(aes(fill = fills),
color = col, size = lwd, linetype = lty)
}
else {
p <- ggplot(t2, aes(dow, woy + 1)) + geom_tile(aes(fill = fills),
fill = low.col, color = col, size = lwd, linetype = lty)
}
if (is.null(from) & is.null(to)) {
weeklabels <- 1:53
if (length(t2$date) == 365) {
weeklabels <- 1:53
}
else {
if (t2$dow[1] == 6) {
weeklabels <- 1:54
}
}
}
else {
weeklabels <- unique(t2$woy) + 1
}
if (is.character(special.days) & wend & length(unique(special.days) ==
length(dates))) {
p <- p + scale_fill_manual(values = special.col,
labels = levels(as.factor(fills)), na.value = "white",
na.translate = FALSE)
}
else {
p <- p + scale_fill_gradient(low = low.col, high = special.col,
na.value = "white")
}
p <- p + facet_wrap(~monlabel, ncol = ncol, scales = "free") +
ggtitle(title) + labs(subtitle = subtitle) + scale_x_continuous(expand = c(0.01,
0.01), position = "top", breaks = seq(0, 6), labels = weekdays) +
scale_y_continuous(expand = c(0.01, 0.01), trans = "reverse",
breaks = unique(t2$woy) + 1, labels = weeklabels) +
geom_text(data = t2, aes(label = gsub("^0+", "",
format(date, "%d"))), size = day.size, family = font.family,
color = days.col, fontface = font.style) + labs(fill = legend.title) +
theme(panel.background = element_rect(fill = NA,
color = NA), strip.background = element_rect(fill = mbg.col,
color = mbg.col), plot.background = element_rect(fill = bg.col),
panel.grid = element_line(colour = ifelse(bg.img ==
"", bg.col, "transparent")), strip.text.x = element_text(hjust = months.pos,
face = font.style, color = months.col, size = months.size),
legend.title = element_text(), axis.ticks = element_blank(),
axis.title = element_blank(), axis.text.y = element_text(colour = week.number.col,
size = week.number.size), axis.text.x = element_text(colour = weeknames.col,
size = weeknames.size * 2.25), plot.title = element_text(hjust = 0.5,
size = title.size, colour = title.col), plot.subtitle = element_text(hjust = 0.5,
face = "italic", colour = subtitle.col, size = subtitle.size),
legend.position = legend.pos, plot.margin = unit(c(1 *
margin, 0.5 * margin, 1 * margin, 0.5 * margin),
"cm"), text = element_text(family = font.family,
face = font.style), strip.placement = "outsite")
if (bg.img != "") {
p <- ggbackground(p, bg.img)
}
}
else {
tidymoons <- data.frame(x = t2$dow + 0.35, y = t2$y +
0.3, ratio = moon, right = right)
tidymoons2 <- data.frame(x = t2$dow + 0.35, y = t2$y +
0.3, ratio = 1 - moon, right = !right)
p <- ggplot(t2, aes(dow, y)) + geom_tile(aes(fill = fills),
color = col, size = lwd, linetype = lty)
if (lunar == TRUE) {
p <- p + geom_moon(data = tidymoons, aes(x, y, ratio = ratio,
right = right), size = lunar.size, fill = "white") +
geom_moon(data = tidymoons2, aes(x, y, ratio = ratio,
right = right), size = lunar.size, fill = lunar.col)
}
if (is.character(special.days) & wend & length(unique(special.days) ==
length(dates))) {
p <- p + scale_fill_manual(values = special.col,
labels = levels(as.factor(fills)), na.value = "white",
na.translate = FALSE)
}
else {
p <- p + scale_fill_gradient(low = low.col, high = special.col,
na.value = "white")
}
p <- p + ggtitle(title) + labs(subtitle = subtitle) +
geom_text(data = df, aes(label = week, x = pos.x,
y = pos.y), size = weeknames.size, family = font.family,
color = weeknames.col, fontface = font.style) +
geom_text(aes(label = texts), color = text.col, size = text.size,
family = font.family) + scale_y_continuous(expand = c(0.05,
0.05), labels = rev(unique(t2$woy)), breaks = 1:length(unique(t2$woy))) +
geom_text(data = t2, aes(label = 1:nrow(filler),
x = dow - 0.4, y = y + 0.35), size = day.size,
family = font.family, color = days.col, fontface = font.style) +
labs(fill = legend.title) + theme(panel.background = element_rect(fill = NA,
color = NA), strip.background = element_rect(fill = NA,
color = NA), plot.background = element_rect(fill = bg.col),
panel.grid = element_line(colour = ifelse(bg.img ==
"", bg.col, "transparent")), strip.text.x = element_text(hjust = 0,
face = "bold", size = months.size), legend.title = element_text(),
axis.ticks = element_blank(), axis.title = element_blank(),
axis.text.y = element_text(colour = week.number.col,
size = week.number.size), axis.text.x = element_blank(),
plot.title = element_text(hjust = 0.5, size = title.size,
colour = title.col), plot.subtitle = element_text(hjust = 0.5,
face = "italic", colour = subtitle.col, size = subtitle.size),
legend.position = legend.pos, plot.margin = unit(c(1 *
margin, 0.5 * margin, 1 * margin, 0.5 * margin),
"cm"), text = element_text(family = font.family,
face = font.style), strip.placement = "outsite")
if (bg.img != "") {
p <- ggbackground(p, bg.img)
}
}
if (pdf == FALSE & doc_name != "") {
warning("Set pdf = TRUE to save the current calendar")
}
if (pdf == TRUE) {
switch(papersize, A6 = {
a <- 148
b <- 105
}, A5 = {
a <- 210
b <- 148
}, A4 = {
a <- 297
b <- 210
}, A3 = {
a <- 420
b <- 297
}, A2 = {
a <- 594
b <- 420
}, A1 = {
a <- 841
b <- 594
}, A0 = {
a <- 1189
b <- 841
}, )
if (doc_name == "") {
if (!is.null(month)) {
doc_name <- paste0("Calendar_", tolower(t2$month[1]),
"_", year, ".pdf")
}
else {
if (!is.null(from) & !is.null(to)) {
doc_name <- paste0("Calendar_", from, "_",
to, ".pdf")
}
else {
doc_name <- paste0("Calendar_", year, ".pdf")
}
}
}
else {
doc_name <- paste0(doc_name, ".pdf")
}
if (orientation == "landscape" | orientation == "l") {
ggsave(filename = if (!file.exists(doc_name))
doc_name
else stop("File does already exist!"), height = b,
width = a, units = "mm")
}
else {
ggsave(filename = if (!file.exists(doc_name))
doc_name
else stop("File does already exist!"), width = b,
height = a, units = "mm")
}
}
return(p)
}
<bytecode: 0xa87c7ef58>
<environment: namespace:calendR>Beware false savings!
Sometimes
“saved” lines of code
turn into
confusing lists of plot specific arguments
Summary
Functions in R
- How to write valid functions
- How to test functions
- How to break down data analysis workflows into smaller tasks (or functions)
- Style considerations for function names, errors and interfaces
- Strategies for designing better functions
Strategies for DRRY functions
Outside in:
Name: What am I trying to do? What evokes the action
Inputs: What pieces of information do I need to provide?
Output: What are we returning?Inside out:
Copy text into body
Identify complexity to manage
Abstract the complexity